Manually measuring and collecting high-quality phenotypic data on crops is painstaking and time-consuming, but necessary for connecting genes to their expressed traits. While high-throughput genomic sequencing is widely available, methods for collecting phenotypic data lag. PSI researchers are developing automated methods for analyzing phenotypes, further advancing researchers’ ability to improve food crops.
Off‐the‐shelf image analysis models outperform human visual assessment in identifying genes controlling seed color variation in sorghum
Seed color in grains is an important phenotype linked to nutritional value and consumer acceptance of new varieties. In sorghum, seed color varies and can indicate the presence of specialized colored metabolites. Typically, it is quantified by qualitative human assessment or biochemical assay. New image-based approaches are potentially more quantitative than human assessment and less expensive than biochemical testing.
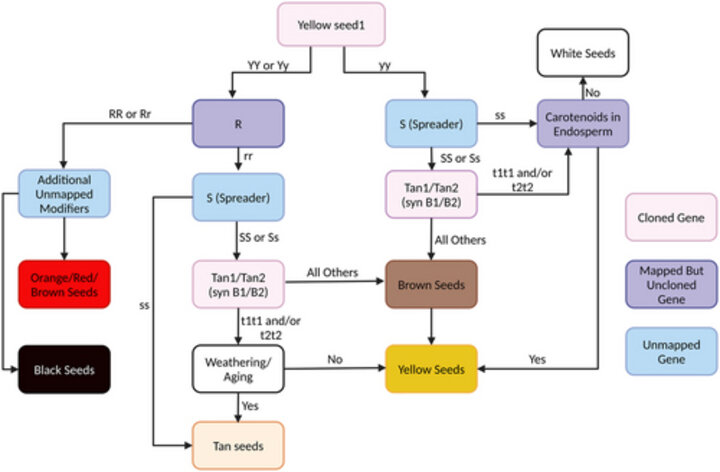
PSI professor James Schnable and his lab tested the feasibility of using image analysis tools trained on rice and wheat to quantify seed color in sorghum using a dataset of over 1,500 images. They found that the measurements of seed color from images were more consistent than human assessment. Genome-wide association studies conducted using imaging data identified more signals near known color genes in sorghum with better support than manually scored seed color for the same experiment. They found the image analysis tools transferable to other species without retraining, which may help researchers develop higher value varieties in sorghum and other grain crops.
Read the article in The Plant Phenome Journal.
Plot‐level satellite imagery can substitute for UAVs in assessing maize phenotypes across multistate field trials
Maize is a critical global crop grown on six continents, and breeding higher-yielding varieties depends on replicated field trials in different environments. While capturing in-season phenotypic and yield data can help accelerate the development of hybrids, collecting these data can be labor intensive and impractical in remote locations.
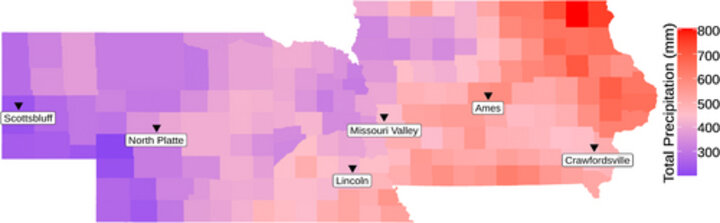
PSI professor James Schable, his lab, and collaborators from several departments at Iowa State University, investigated the use of satellite remote sensing in crop performance evaluation. The team generated a dataset of over 20,000 images of over 80 maize hybrids in the US corn belt, collected by satellite and drones and corresponding actual yield measurements. They discovered that predictive models using satellite data for forecasting yields performed as well as models using drone data, opening a path to further approaches to predicting plant parameters from satellite images and environmental data.
Read the article in Plants, People, Planet.